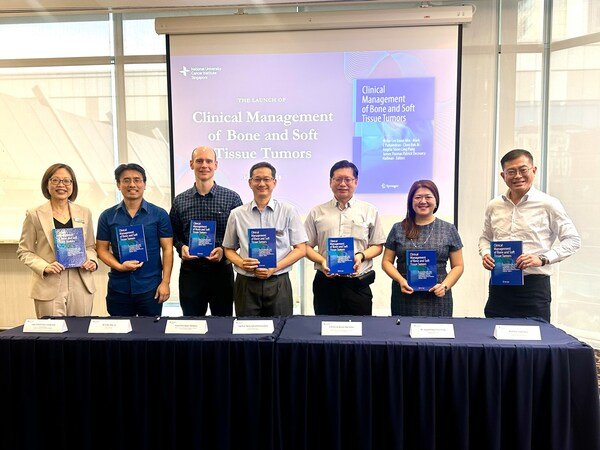
Cellworks Therapy Biosimulation Study Reveals Promising New Biomarkers for TMZ Resistance in Glioblastoma Patients
23 November 2022 | Wednesday | News

Image Source : Public Domain
Cellworks Group Inc., a leader in Precision Drug Development and Personalized Therapy Biosimulation, today announced results from a Cellworks biosimulation pilot study, which examined the impact of mismatch repair deficiency (MRD) on survival of temozolomide (TMZ)-treated patients with MGMT methylated (m-MGMT) glioblastoma (GBM). The biosimulation study validated that TMZ does not trigger apoptosis in mismatch repair deficiency cancers and uncovered promising new biomarkers for TMZ resistance. The study also found inferior overall survival (OS) for MRD compared to the historical experience of unmethylated-MGMT patients, suggesting that TMZ-induced hypermutation may compromise survival.
Results from the study were featured in a poster presentation at the 27th Annual Meeting and Education Day of the Society for Neuro-Oncology on November 18, 2022 and available online at Neuro-Oncology Journal Article EPCO-06.
“It has been a surprise to most oncologists that DNA mismatch repair (MMR) deficiency determines temozolomide success against GBM and that MMR is much more complicated than merely mutation or deletion of the four MMR genes,” said Dr. Michael Castro M.D., Neuro-oncologist, Medical Oncologist at Beverly Hills Cancer Center; Chief Medical Officer at Cellworks, Group Inc. and Principal Investigator of the study. “In fact, Cellworks’ biosimulation identifies MMR deficiency due to epigenetic, translational and proteomic silencing that leads to early progression on temozolomide despite methylated MGMT. To a significant extent, these other mechanisms of MMR deficiency explain the long observed anomaly that many patients expected to respond to temozolomide based on methylated MGMT have no chance to benefit. Crucially, they could be treated instead with lomustine which does not require MMR for efficacy. Hopefully, biosimulation of MMR deficiency should emerge as an important new biomarker in neurooncology that allows physicians to make more effective treatment decisions in the front-line management of GBM.”
Cellworks Therapy Biosimulation
The Cellworks Platform biosimulates how a patient's personalized genomic disease model will respond to therapies prior to treatment and can also identify novel drug combinations for treatment-refractory patients. The platform is powered by the groundbreaking Cellworks Computational Biology Model (CBM), a highly curated mechanistic network of 6,000+ human genes, 30,000 molecular species and 600,000+ molecular interactions. As part of the biosimulation process, personalized disease models are created for each patient using their cytogenetic and molecular data as input to the Cellworks CBM. The Cellworks platform analyzes the impact of specific therapies on the patient’s personalized disease model and predicts the efficacy of specific chemotherapies.
Biosimulation Study
The purpose of the study was to examine the impact of mismatch repair deficiency (MRD) on survival of temozolomide (TMZ)-treated patients with MGMT methylated (m-MGMT) glioblastoma (GBM) using Cellworks computational biosimulation.
Background
TMZ-induced G:T mismatches trigger mismatch repair (MMR) to perform futile repair of O6-meG necessary to achieve apoptosis. Without intact MMR, the G:T mutation does not cause cell death, but instead is genomically incorporated to produce DNA mutation signature #11. Hypermutation ensues and may compromise survival without the redemptive benefit of immunotherapy. MMR genes are infrequently mutated or deleted in GBM. More commonly, MRD results from epigenetic, translational and proteomic silencing.
Methods
Comprehensive genomic profiling and Cellworks biosimulation was utilized to diagnose MRD and correlated with survival in 38 TCGA patients with newly diagnosed IDH wildtype, m-MGMT GBM treated with adjuvant TMZ. The integrity of MMR (MutSa and MutLa) was biosimulated for each patient using a computational model of signaling pathway behavior. Kaplan-Meier curves were constructed for progression-free survival (PFS) and overall survival (OS) using the treatment start date for calculation.
Results
Biosimulation results indicate that TMZ does not trigger apoptosis in mismatch repair deficiency cancers. Overall Survival (OS) in MRD patients was worse than historical u-MGMT patients who benefit little from TMZ. MMR had higher MMR biosimulation scores compared to patients with one or more abnormalities resulting in MRD (p = 0.00082). Half of MRD patients had classical COSMIC MMR mutation signatures representing a portion of the mutations in the tumor. Five-year survivors were only found in the MMR proficient group. Partial MRD from epigenetic silencing can result in MMR mutation signatures, but did not cause hypermutation and MSI.
Nineteen of the 38 patients with MMR deficiency had 1 to 9 pathways compromised, impacting MMR function. Six patients had deletions of MLH1, HSH2 or MSH6. Other abnormalities included loss of EP300, CREBBP, KMT2A-D, ARID1A, HUS or EXO1. One of the 38 patients had MSI from germline MRD. Hypermutation was not identified in 37 of the 38 patients.
Conclusions
This study shows that biosimulation of MRD can identify m-MGMT patients who develop early progression on TMZ, echoing the long-established knowledge that TMZ does not trigger apoptosis in MRD cancers. Lomustine does not rely on intact MMR for efficacy and therefore it may blunt the impact of MMR on OS or could produce better disease control if used as an initial therapy. The observed deleterious impact of MRD on survival for TMZ treated m-MGMT patients should be examined and confirmed in a larger patient cohort.
Most Read
- How Does GLP-1 Work?
- Innovations In Magnetic Resonance Imaging Introduced By United Imaging
- Management of Relapsed/Refractory Multiple Myeloma
- 2025 Drug Approvals, Decoded: What Every Biopharma Leader Needs to Know
- BioPharma Manufacturing Resilience: Lessons From Capacity Expansion and Supply Chain Resets from 2025
- APAC Biopharma Review 2025: Innovation, Investment, and Influence on the Global Stage
- Top 25 Biotech Innovations Redefining Health And Planet In 2025
- The New AI Gold Rush: Western Pharma’s Billion-Dollar Bet on Chinese Biotech
- Single-Use Systems Are Rewiring Biopharma Manufacturing
- The State of Biotech and Life Science Jobs in Asia Pacific – 2025
- Asia-Pacific Leads the Charge: Latest Global BioSupplier Technologies of 2025
- Invisible Threats, Visible Risks: How the Nitrosamine Crisis Reshaped Asia’s Pharmaceutical Quality Landscape
Bio Jobs
- Sanofi Turns The Page As Belén Garijo Steps In And Paul Hudson Steps Out
- Global Survey Reveals Nearly 40% of Employees Facing Fertility Challenges Consider Leaving Their Jobs
- BioMed X and AbbVie Begin Global Search for Bold Neuroscience Talent To Decode the Biology of Anhedonia
- Thermo Fisher Expands Bengaluru R&D Centre to Advance Antibody Innovation and Strengthen India’s Life Sciences Ecosystem
- Accord Plasma (Intas Group) Acquires Prothya Biosolutions to Expand Global Plasma Capabilities
- ACG Announces $200 Million Investment to Establish First U.S. Capsule Manufacturing Facility in Atlanta
- AstraZeneca Invests $4.5 Billion to Build Advanced Manufacturing Facility in Virginia, Expanding U.S. Medicine Production
News
Editor Picks