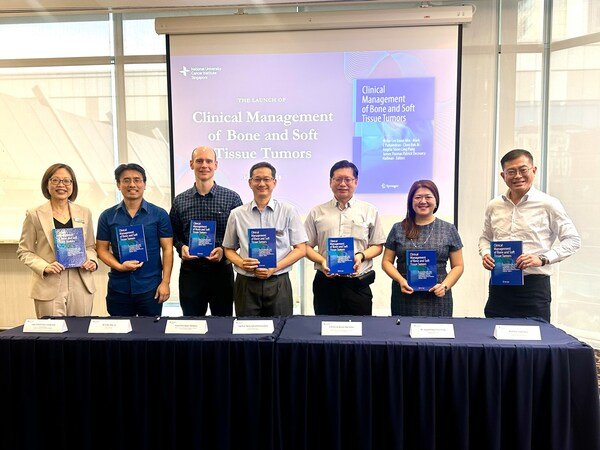
Simplifying Drug Discovery: How Molecule AI's Empowers Researchers - An Interview with Molecule AI's Saurabh Singal
12 October 2023 | Thursday | News

In an illuminating interview with BioPharma APAC, Saurabh Singal, the CEO-Founder of Molecule AI, delves into the transformative power of Molecule AI's platform, Molecule GEN, and how it is reshaping the landscape of in-silico computational drug discovery. Singal discusses the unique features that set Molecule GEN apart in this field, such as its comprehensive approach addressing diverse factors in drug discovery. He also explains how Molecule GEN streamlines complex manual tasks for professionals in biotechnology, chemistry, and AI research, empowering them with automation and a wealth of valuable information. Furthermore, Singal shares success stories that highlight how Molecule GEN significantly reduces time and effort in candidate identification and validation, ultimately accelerating the drug development process. The interview also sheds light on the decision to offer Molecule GEN as a Software as a Service (SaaS) platform, making cutting-edge AI accessible while simplifying the complexities of computational resources. Finally, Singal reveals how Molecule AI's capacity to explore uncharted territories in drug discovery opens the door to pioneering candidates and showcases recent advancements and updates that position researchers at the forefront of technological innovation in this vital field. This interview provides a comprehensive look into the future of drug discovery and the critical role that Molecule AI plays in shaping it.
Can you provide more insight into how Molecule GEN's AI-driven platform is transforming the landscape of in-silico computational drug discovery? What specific features or capabilities make it stand out in the field?
While there are several powerful tools already in the market for computational drug discovery, existing solutions do not address the totality of factors such as molecule types (e.g. proteins and small molecules), target awareness, Absorption-Distribution-Metabolism-Excretion (ADME) properties, Toxicity screening, ease of use, integration of different parts of the drug discovery pipeline, etc. Our platform addresses all of these factors and provides widely applicable, easy-to-use, modules for all levels of users.
Our platform enables a wide range of workflows including target identification, analysis of the structure and properties of the target vis-a-vis related proteins, identification of active sites, de-novo generation of molecules (both small molecules and antibodies) against the target, prediction and optimization of ADMET properties of the generated molecules, docking and molecular dynamics. Many of these workflows incorporate powerful in-house AI algorithms which are improvements over current state-of-the-art.
We believe that such a comprehensive package of workflows within a single platform can be a powerful enabler for success for researchers in computational drug discovery, and is a unique feature of our platform.
Molecule GEN is described as a platform that empowers researchers by automating manual tasks and providing a wealth of information. Could you elaborate on some examples of these tasks and how the platform simplifies them for professionals in biotechnology, chemistry, and AI research?
As an example of the kind of convenient automation our platform can provide, consider the scenario where a researcher has a protein target in mind. Let’s say this target was the N-methyl D-aspartate (NMDA) receptor, a protein implicated in many neurological conditions. In order to discover drugs against this target, the researcher needs to look at the structure of the target. Such structures could typically be obtained from experimental data submitted in the protein data bank (PDB) server. However, a search for “NMDA receptor” in the PDB returns tens of thousands of results, and though the results are ranked by a relevance metric, it is not a trivial matter for the researcher to assess which of these structures is the most relevant in their particular context. Importing any of these PDB structures into Molecule GEN allows the researcher to access a wealth of metadata in a convenient form that enables the assessment of the right structure to be used. For example, a much shorter list of all related PDB structures, clustered according to multiple similarity criteria, are provided along with the imported structure. For each of these structures, a host of information‒including experimental conditions, quality of the structure, number and nature of protein units, the length of amino acid sequence covered, presence of various cofactors, ligands, and inhibitors‒are given. This places each structure in the right context and makes it much easier for the researcher to decide which one(s) to continue working with.
As another example, consider the scenario where the researcher would like to narrow down a set of molecules based on physicochemical or ADMET properties. Molecule GEN offers filters based on multiple simultaneous and customizable criteria, which can be implemented in a single click, resulting in a smaller set of lead molecules.
Efficiency in drug discovery is a critical aspect of Molecule GEN. Could you share some success stories or case studies where the platform has significantly reduced the time and effort required for candidate identification and validation? How does this impact the drug development process?
We are testing and optimizing the Molecule GEN workflow by generating lead-like molecules against chosen targets. With the workflow as it currently stands, we believe that given a target, we can generate easily synthesizable lead compounds with the right drug-like properties in about two weeks (given the computational resources of a single A100 GPU). That is an order of magnitude shortening of timelines as compared to the corresponding experimental process. Using this quick workflow, we have generated molecules against CNS targets, and tyrosine kinase targets.
Even when considering a scenario where a set of molecules already exists, our models for prediction of ADMET and physicochemical properties can help in creating an order of priority for individual molecules in the set. As an example, when we apply our cardiotoxicity prediction models to approved drugs with publicly available cardiotoxicity data (“Yes” or “No”, as per the standard hERG blockade assay), we see that molecules predicted as being toxic have an 85% probability of actually being toxic.
This kind of predictive ability ensures that the molecules being advanced as leads from our pipeline are the right ones that are likely to succeed in experimental testing, thus cutting down time and cost for getting to the molecule that will succeed in the clinic.
The decision to offer Molecule GEN as a Software as a Service (SaaS) platform is interesting. Can you explain the advantages of this approach for researchers, especially in terms of accessibility and simplifying complex technicalities related to computational resources?
The process of setting up the software stack for end to end drug discovery is a challenging task. With the advent of machine learning models, the application software needs powerful GPU clusters to run the pipelines. The optimization of the software to work at highest possible throughput is achieved through careful tuning of multiple parameters. Molecule GEN does all this heavy lifting behind the scenes and offers the best returns for the hardware cost. The users eliminate the process of procuring, setting up, optimizing and maintaining the complex computational resources required, and can run Molecule GEN from any web browser. With the complexities of hardware and software resources out of their way, users can instead focus on drug discovery, thus removing a significant hurdle for many drug discovery scientists in adopting cutting edge AI methods in their research.
Molecule AI's capacity to explore uncharted territories in drug discovery is intriguing. How does the platform provide insights into novel chemical spaces, and what are the potential implications of this for researchers in terms of discovering groundbreaking candidates?
Traditional drug discovery methods rely on screening existing molecular datasets, which, despite being extensive, represent only a minuscule fraction of the entire possible molecular space. Molecule GEN leverages Generative AI to intelligently explore the vast molecular and antibody-sequence spaces beyond the limits of traditional datasets. By learning patterns and relationships between molecular structures and their interactions with target proteins, it can predict potential candidates without the need for exhaustive screenings. It can generate entirely new molecular structures or antibody sequences that might not have been thought of or synthesized yet. For researchers, this means access to groundbreaking molecules and antibodies that might have been overlooked with conventional methods, and facilitates a leap from iterative, incremental improvements to groundbreaking discoveries.
A difficulty in actually achieving this is the lack of experimental data to train models to learn the “right” and “wrong” structural features of antibodies against specific targets. At Molecule AI, we have devised ways of generating high-quality training data through advanced simulation methods.
Molecule AI is continuously evolving with state-of-the-art generative models for drug design. Can you highlight some recent advancements or updates to the platform, and how these contribute to keeping researchers at the forefront of technological innovation in drug discovery?
At Molecule AI, we are steadfast in our commitment to harnessing cutting-edge generative models for drug discovery. Recently, we've integrated advanced generative AI models tailored for a spectrum of drug design applications, spanning from ligand and antibody design to molecular docking. Additionally, we're pioneering robust neural modules dedicated to ensuring the safety of the molecules that are designed. These enhancements position researchers who will use the platform at the vanguard of technological innovation in the drug discovery landscape.
Most Read
- How Does GLP-1 Work?
- Innovations In Magnetic Resonance Imaging Introduced By United Imaging
- Management of Relapsed/Refractory Multiple Myeloma
- 2025 Drug Approvals, Decoded: What Every Biopharma Leader Needs to Know
- BioPharma Manufacturing Resilience: Lessons From Capacity Expansion and Supply Chain Resets from 2025
- APAC Biopharma Review 2025: Innovation, Investment, and Influence on the Global Stage
- Top 25 Biotech Innovations Redefining Health And Planet In 2025
- The New AI Gold Rush: Western Pharma’s Billion-Dollar Bet on Chinese Biotech
- Single-Use Systems Are Rewiring Biopharma Manufacturing
- The State of Biotech and Life Science Jobs in Asia Pacific – 2025
- Asia-Pacific Leads the Charge: Latest Global BioSupplier Technologies of 2025
- Invisible Threats, Visible Risks: How the Nitrosamine Crisis Reshaped Asia’s Pharmaceutical Quality Landscape
Bio Jobs
- Sanofi Turns The Page As Belén Garijo Steps In And Paul Hudson Steps Out
- Global Survey Reveals Nearly 40% of Employees Facing Fertility Challenges Consider Leaving Their Jobs
- BioMed X and AbbVie Begin Global Search for Bold Neuroscience Talent To Decode the Biology of Anhedonia
- Thermo Fisher Expands Bengaluru R&D Centre to Advance Antibody Innovation and Strengthen India’s Life Sciences Ecosystem
- Accord Plasma (Intas Group) Acquires Prothya Biosolutions to Expand Global Plasma Capabilities
- ACG Announces $200 Million Investment to Establish First U.S. Capsule Manufacturing Facility in Atlanta
- AstraZeneca Invests $4.5 Billion to Build Advanced Manufacturing Facility in Virginia, Expanding U.S. Medicine Production
News
Editor Picks