Scientists discover and name novel gene that governs left-right asymmetry within the human body
14 December 2021 | Tuesday | News

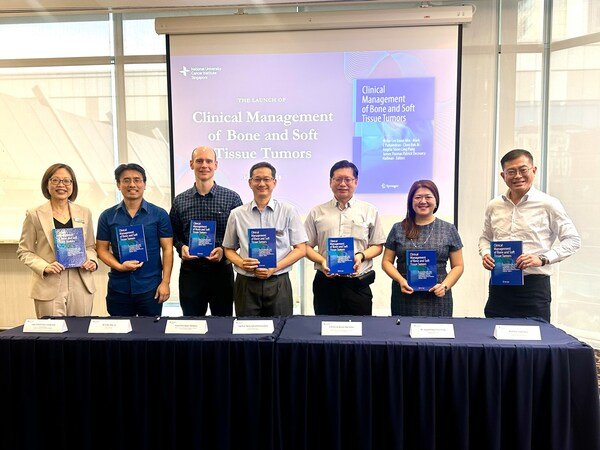
Chest X-ray of a control individual (left), with his heart pointing to the left (L), while the chest X-ray of a patient with heterotaxy (right) revealed that his heart is inversely positioned pointing instead to the right (R).
A team of researchers, led by the Agency for Science, Technology and Research’s (A*STAR) Genome Institute of Singapore (GIS), in collaboration with A*STAR’s Institute of Molecular and Cell Biology (IMCB) and Bioinformatics Institute (BII), and clinicians from six different countries, has discovered a novel gene, Ciliated Left-Right Organizer Metallopeptidase (CIROP), that is crucial for establishing proper left-right asymmetry during vertebrate embryonic development. Babies that carry mutations in CIROP had internal organs randomly positioned, leading to severe birth defects consistent with heterotaxy
At first glance, the human body looks symmetrical because our left side appears to be a mirror image of the right side. This symmetry is in fact only skin-deep since our internal organs are asymmetrically placed – the heart and spleen are on the left side while the liver is on the right side of our body. This is governed by a set of genes – comprising CIROP, PKD1L1, MMP21, DAND5 and C1orf127 – which act early during embryonic development to assign each organ a stereotypical position. When this is not achieved properly, babies can be born with birth anomalies such as congenital heart defects and misplacement of internal organs along the left-right axis. On average, these diseases occur once every 10,000 births and are grouped under syndromes of heterotaxy.
By performing an evolutionary analysis of genomes from many vertebrate species, CIROP, PKD1L1, MMP21, DAND5 and C1orf127 were found to be present in ancestral animals such as fish and frogs, but absent in reptiles, birds, and certain mammals such as cetaceans. This pattern of gene disappearance during evolution correlates with the loss of motile cilia in the transient organ that establishes left-right patterning during embryogenesis.
Prof Bruno Reversade, Senior Group Leader of the Laboratory of Human Genetics and Therapeutics at GIS and IMCB, said, “Our phylogenetic screen for genes that have disappeared in vertebrate species yielded important evolutionary insights into the development of left-right patterning. Our findings suggest that these five genes have only one function, which is to distinguish left from right. Active during a small window in the course of development, these genes may never be used again after birth.”
Dr Emmanuelle Szenker-Ravi, Research Scientist from the Laboratory of Human Genetics and Therapeutics at GIS, and first author of this study, was struck that CIROP, which is such an essential gene, had never been characterised before, “CIROP’s incomplete annotation in the human genome prevented it from being used for diagnostic purposes via exome sequencing. We are thrilled that this is now remedied and will immediately benefit affected families.”
Prof Patrick Tan, Executive Director of GIS, said, “The study illustrates the power of Mendelian genetics2 which assigns gene functions and provides indisputable causality. Birth defects due to genetic mutations have devastating impact on both the children and their families. These findings will serve as the basis for research on potential therapies.”
Most Read
- How Does GLP-1 Work?
- Innovations In Magnetic Resonance Imaging Introduced By United Imaging
- Management of Relapsed/Refractory Multiple Myeloma
- 2025 Drug Approvals, Decoded: What Every Biopharma Leader Needs to Know
- BioPharma Manufacturing Resilience: Lessons From Capacity Expansion and Supply Chain Resets from 2025
- APAC Biopharma Review 2025: Innovation, Investment, and Influence on the Global Stage
- Top 25 Biotech Innovations Redefining Health And Planet In 2025
- The New AI Gold Rush: Western Pharma’s Billion-Dollar Bet on Chinese Biotech
- Single-Use Systems Are Rewiring Biopharma Manufacturing
- The State of Biotech and Life Science Jobs in Asia Pacific – 2025
- Asia-Pacific Leads the Charge: Latest Global BioSupplier Technologies of 2025
- Invisible Threats, Visible Risks: How the Nitrosamine Crisis Reshaped Asia’s Pharmaceutical Quality Landscape
Bio Jobs
- Sanofi Turns The Page As Belén Garijo Steps In And Paul Hudson Steps Out
- Global Survey Reveals Nearly 40% of Employees Facing Fertility Challenges Consider Leaving Their Jobs
- BioMed X and AbbVie Begin Global Search for Bold Neuroscience Talent To Decode the Biology of Anhedonia
- Thermo Fisher Expands Bengaluru R&D Centre to Advance Antibody Innovation and Strengthen India’s Life Sciences Ecosystem
- Accord Plasma (Intas Group) Acquires Prothya Biosolutions to Expand Global Plasma Capabilities
- ACG Announces $200 Million Investment to Establish First U.S. Capsule Manufacturing Facility in Atlanta
- AstraZeneca Invests $4.5 Billion to Build Advanced Manufacturing Facility in Virginia, Expanding U.S. Medicine Production
News
Editor Picks